The new Illumina patterned flow cell technology uses chemistry that is prone to “index hopping”, leading to reads being assigned to the wrong sample in multiplexed sequencing runs. This problem has already caused countless research groups to re-assess their experimental results or refrain from using the new sequencers. However, with careful experimental preparation it may be possible to ameliorate this situation so as to become negligible for most applications
June 7, 2017
Steven Wingett
HiSeq, Illumina, X Ten, All Applications
Want to keep up to date with new articles?
We have a
mailing list and
twitter feed to help with that..
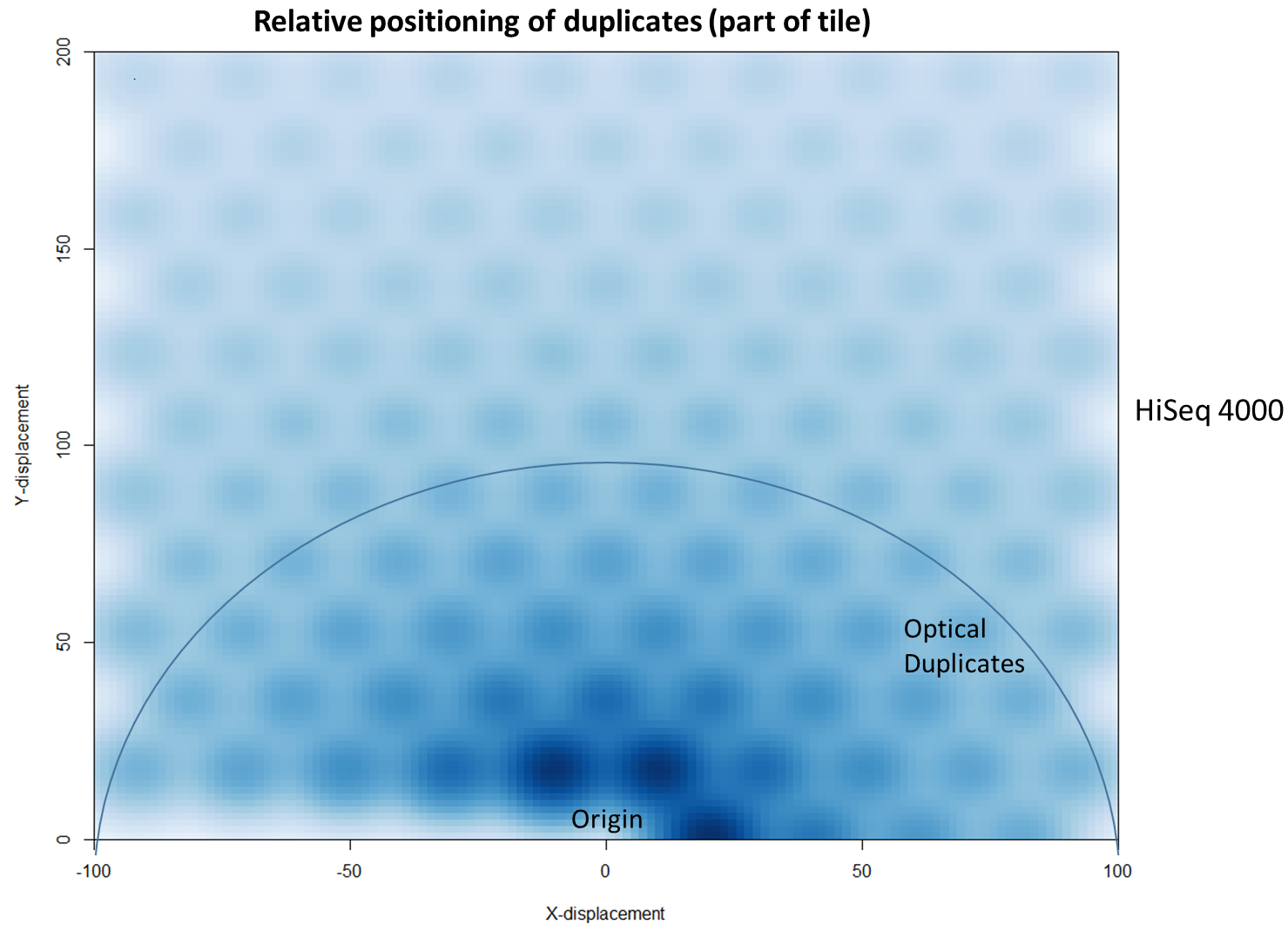
The latest Illumina sequencers – such as the HiSeq X, HiSeq 3000 and HiSeq 4000 – use patterned flow cells to enable the discrimination between much more densely packed DNA clusters. While such technology substantially increases the number of reads generated per sequence run, this innovation may lead to an increased number of duplicates, thereby negating the improved yield and making subsequent data analysis potentially more difficult. Further investigation shows that these putative sequencing duplicates are generally in close two-dimensional proximity on a flow cell, which may provide an opportunity to develop bioinformatics solutions to identify and discard such artefacts.
March 2, 2017
Steven Wingett
HiSeq, All Applications, Bowtie2, HiCUP, Picard
Soft-clipping of sequencing reads allows the masking of portions of the reads that do not align to the genome from end to end, which may be desirable for certain types of analysis (e.g. detection of structural variants). For standard alignment processes soft-clipping may however incorrectly trim reads and lead to the mis-assignments of reads primarily to repetitive regions. This phenomenon appears to vary in severity for different sequencing applications with Bisulfite sequencing being worst off.
May 16, 2016
Felix Krueger
All Applications, Bismark, Bowtie2, bwa-meth, HISAT2, SeqMonk, Trim Galore!
With the introduction of the NextSeq system Illumina changed the way their image data was acquired so that instead of capturing 4 images per cycle they needed only 2. This speeds up image acquisition significantly but also introduces a problem where high quality calls for G bases can be made where there is actually no signal on the flowcell.
May 4, 2016
Simon Andrews
NextSeq, All Applications, Cutadapt, FastQC
With the increasing capacity of a single flowcell lane it can be tempting to mix samples of different types within the same lane to make the most of your sequencing, but cross contamination between libraries in a flowcell can lead to the generation of artefacts which can mess up your analysis.
April 15, 2016
Simon Andrews
Illumina, All Applications, SeqMonk

Probably the single biggest problem with the mapping of reads to a reference sequence is dealing with reads which come from parts of the genome which aren’t in the assembly. These reads can cause significant amounts of noise in anlayses performed on genomic data.
March 21, 2016
Simon Andrews
All Technologies, All Applications

Paired-end libraries generated by Post Bisulfite Adapter Tagging (PBAT) often suffer from poorer mapping efficiencies when compared to standard whole genome shotgun Bisulfite-Seq libraries. In addition to the usual suspects that have a detrimental impact on mapping efficiency we found that a substantial proportion of paired-end PBAT libraries appears to consist of chimeric reads that map to different places in the genome, not unlike Hi-C type experiments. Chimeric reads also affect single-cell libraries (scBS-seq) as they are constructed using a PBAT approach.
March 18, 2016
Felix Krueger
Illumina, Methylation, PBAT, Bismark, Cutadapt, SeqMonk, Trim Galore!

One of the standard fields in the SAM/BAM file format is the mapping quality (MAPQ) value. This value can be very useful to help filter mapped reads before doing downstream analysis – unfortunately the implementation of this value is in no way consistent between different aligners so it takes a fair bit of research to know how to use it appropriately. Mis-applying the filter could cause reads to be inappropriately excluded from an analysis.
March 17, 2016
Simon Andrews
All Technologies, All Applications, BamQC, SeqMonk
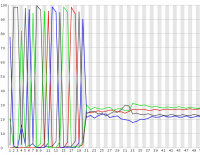
In some experimental designs a large proportion of the sequences in a library can have identical sequence at their 5′ end. These types of library can cause problems for the data collection and base calling on illumina sequencers, leading to the generation of poor quality data.
March 15, 2016
Simon Andrews
Illumina, All Applications, FastQC
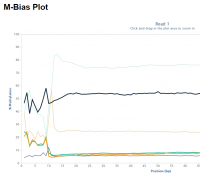
Random priming in PBAT libraries introduces drastic biases in the base composition and methylation levels especially at the 5′ end of all reads. As a result, affected bases should be removed from the libraries before the alignment step.
March 11, 2016
Felix Krueger
Illumina, Methylation, PBAT, BamQC, Bismark, FastQC, Trim Galore!
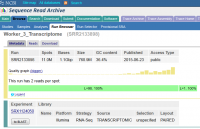
We are increasingly re-using data deposited in public sequence archives such as SRA, ENA or DDBJ and we rely on being able to successfully extract data from these sources. In some cases we have found that errors in the validation of the data can mean that data is corrupted when it is downloaded from these repositories.
March 3, 2016
Simon Andrews
All Technologies, All Applications
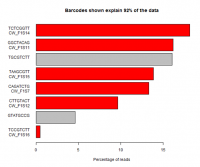
When running multiple samples in the same sequencing reaction the different libraries are usually created with unique sequence tags (barcodes) to allow the sequence from the lane to be separated. Problems during this splitting are common and can have serious effects on downstream analysis.
February 12, 2016
Simon Andrews
All Technologies, All Applications, Data Processing

Library construction of standard directional BS-Seq samples often consist of several steps including sonication, end-repair, A-tailing and adapter ligation. Since the end-repair step typically uses unmethylated cytosines for the fill-in reaction the filled-in bases will generally appear unmethylated after bisulfite conversion irrespective of their true genomic methylation state.
February 12, 2016
Felix Krueger
Illumina, BS-Seq, Methylation, Bismark, Data Processing
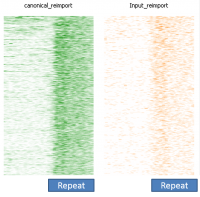
In a dataset where you have some degree of technical duplication, and have not filtered your data to only keep uniquely mapping reads then if you perform deduplication it will look as if repeat sequences are enriched.
February 11, 2016
Simon Andrews
ChIP-Seq, Data Processing, SeqMonk
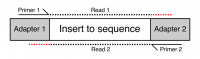
Many sequencing platforms require the addition of specific adapter sequences to the end of the fragments to be sequenced. For an individual fragment, if the length of the sequencing read is longer than the fragment to be sequenced then the read will continue into the adapter sequence on the end. Unless it is removed this adapter sequence will cause problems for downstream mapping, assembly or other analysis.
February 7, 2016
Simon Andrews
Cutadapt, FastQC, Skewer, Trim Galore!

The assumption when analysing sequence datasets is that every sequence comes from a different biological fragment in the original sample. Many library preparation techniques though include one or more PCR steps which introduce the possibility that the same original fragment can be observed multiple times, biasing the results produced. In some cases this type of duplication can be extreme and have a serious effect on the ability to analyse the data correctly.
February 6, 2016
Simon Andrews
FastQC, MultiQC, Preseq, SeqMonk

For speed and efficiency many RNA-Seq mapping protocols map either entirely or initially to a transcriptome sequence. When the sample is contaminated with genomic material you can get significant numbers of reads reported as uniquely mapping when they actually map to many locations within the genome.
February 2, 2016
Simon Andrews
mRNA-Seq, SeqMonk

In theory RNA-Seq samples only contain reads derived from RNA. Most protocols include a DNase treatment to remove any remaining DNA from the sample. It is fairly common however to see samples with significant amounts of DNA contamination. Ignoring this can bias the counts generated from the data and throw off normalisation.
February 2, 2016
Simon Andrews
RNA-Seq, SeqMonk

One of the biggest problems with sequencing libraries is that the material might be contaminated with something unexpected. One of the simplest forms of contamination is where you have material from a different species than expected. In many cases the rogue material will come from a species you can guess based on their other species commonly used in your lab. Screening for this type of contamination will help spot when you have contaminated samples, and can also help when you have completely switched samples.
February 1, 2016
Simon Andrews
FastQ Screen
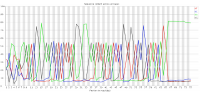

The construction of sequencing libraries on many platforms requires the addition of specific adapter sequences to the ends of the fragments to be sequenced. Although there are steps in place to ensure that only valid adapter+insert combinations make it onto the sequencer it is possible to get adapter dimers with no valid insert making it through the sequencing process.
February 1, 2016
Simon Andrews
Illumina, FastQC





















