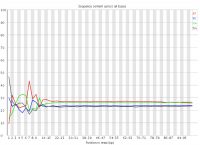
In a randomly primed library there is no reason to expect that a specific sequencing cycle should contain more of one specific base than any other cycle. It is commonly observed though that this type of library does contain a cycle-specific sequence bias, most frequently in the initial bases of the run.
January 31, 2016
Simon Andrews
mRNA-Seq, FastQC
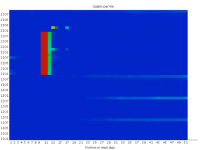

Rather than a general loss of quality for a whole sequencing lane or run sometimes partial failures occur. These can affect specific regions and cycles and can have knock on effects for the data generated
January 31, 2016
Simon Andrews
Illumina, FastQC
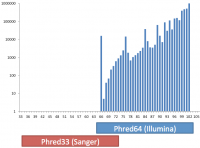
Base call qualities in fastq and BAM format files are recorded using a text based encoding. Mistakes in the way this encoding is applied can lead to incorrect interpretation of base call accuracies in downstream analyses.
January 20, 2016
Simon Andrews
FastQC, QC Software
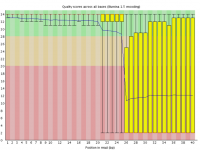
Sometimes a sequencing run can experience a sudden and lasting loss of base call quality across all sequences.
January 20, 2016
Simon Andrews
Illumina, FastQC, QC Software
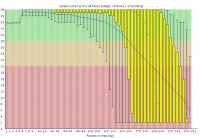
Illumina based sequencing shows a loss of base call quality as the number of sequencing cycles performed increases.
January 20, 2016
Simon Andrews
Illumina, FastQC, QC Software







