
Paired-end libraries generated by Post Bisulfite Adapter Tagging (PBAT) often suffer from poorer mapping efficiencies when compared to standard whole genome shotgun Bisulfite-Seq libraries. In addition to the usual suspects that have a detrimental impact on mapping efficiency we found that a substantial proportion of paired-end PBAT libraries appears to consist of chimeric reads that map to different places in the genome, not unlike Hi-C type experiments. Chimeric reads also affect single-cell libraries (scBS-seq) as they are constructed using a PBAT approach.
March 18, 2016
Felix Krueger
Illumina, Methylation, PBAT, Bismark, Cutadapt, SeqMonk, Trim Galore!
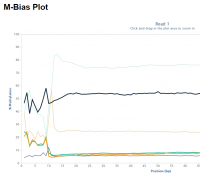
Random priming in PBAT libraries introduces drastic biases in the base composition and methylation levels especially at the 5′ end of all reads. As a result, affected bases should be removed from the libraries before the alignment step.
March 11, 2016
Felix Krueger
Illumina, Methylation, PBAT, BamQC, Bismark, FastQC, Trim Galore!

Library construction of standard directional BS-Seq samples often consist of several steps including sonication, end-repair, A-tailing and adapter ligation. Since the end-repair step typically uses unmethylated cytosines for the fill-in reaction the filled-in bases will generally appear unmethylated after bisulfite conversion irrespective of their true genomic methylation state.
February 12, 2016
Felix Krueger
Illumina, BS-Seq, Methylation, Bismark, Data Processing




